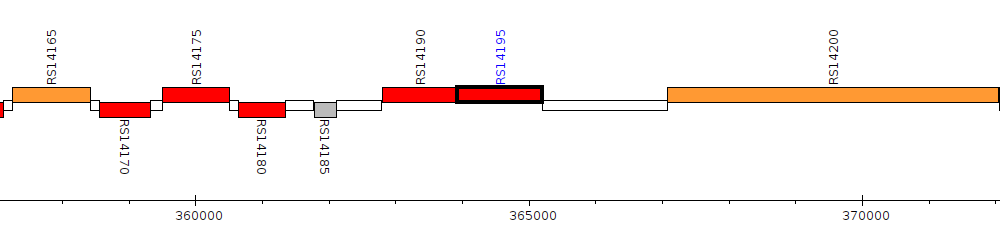
Pseudomonas sp. WCS358, PC358_RS14195
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0000271 | polysaccharide biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF500136
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016628 | oxidoreductase activity, acting on the CH-CH group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF500136
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PF03720
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051287 | NAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF03720
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 5 | 198 | 6.51E-44 |
| SUPERFAMILY | SSF52413 | UDP-glucose/GDP-mannose dehydrogenase C-terminal domain | IPR036220 | UDP-glucose/GDP-mannose dehydrogenase, C-terminal domain superfamily | 314 | 417 | 1.23E-18 |
| NCBIfam | TIGR03026 | JCVI: nucleotide sugar dehydrogenase | IPR017476 | UDP-glucose/GDP-mannose dehydrogenase | 5 | 402 | 1.6E-119 |
| SUPERFAMILY | SSF48179 | 6-phosphogluconate dehydrogenase C-terminal domain-like | IPR008927 | 6-phosphogluconate dehydrogenase-like, C-terminal domain superfamily | 206 | 299 | 4.59E-29 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 1 | 204 | 1.5E-67 |
| FunFam | G3DSA:3.40.50.720:FF:000139 | UDP-N-acetyl-D-mannosamine dehydrogenase | - | - | 1 | 202 | 5.4E-105 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 238 | 420 | 3.2E-47 |
| Pfam | PF00984 | UDP-glucose/GDP-mannose dehydrogenase family, central domain | IPR014026 | UDP-glucose/GDP-mannose dehydrogenase, dimerisation | 206 | 293 | 7.0E-28 |
| PIRSF | PIRSF000124 | UDPglc_GDPman_dh | IPR017476 | UDP-glucose/GDP-mannose dehydrogenase | 4 | 419 | 1.4E-74 |
| Gene3D | G3DSA:1.20.5.100 | - | - | - | 206 | 236 | 4.0E-6 |
| SMART | SM00984 | UDPG_MGDP_dh_C_a_2_a | IPR014027 | UDP-glucose/GDP-mannose dehydrogenase, C-terminal | 325 | 420 | 3.4E-12 |
| PANTHER | PTHR43491 | UDP-N-ACETYL-D-MANNOSAMINE DEHYDROGENASE | IPR028359 | UDP-N-acetyl-D-mannosamine/glucosamine dehydrogenase | 3 | 418 | 0.0 |
| Pfam | PF03720 | UDP-glucose/GDP-mannose dehydrogenase family, UDP binding domain | IPR014027 | UDP-glucose/GDP-mannose dehydrogenase, C-terminal | 325 | 418 | 8.0E-15 |
| Pfam | PF03721 | UDP-glucose/GDP-mannose dehydrogenase family, NAD binding domain | IPR001732 | UDP-glucose/GDP-mannose dehydrogenase, N-terminal | 5 | 189 | 7.6E-59 |
| PIRSF | PIRSF500136 | UDP_ManNAc_DH | IPR028359 | UDP-N-acetyl-D-mannosamine/glucosamine dehydrogenase | 1 | 420 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.