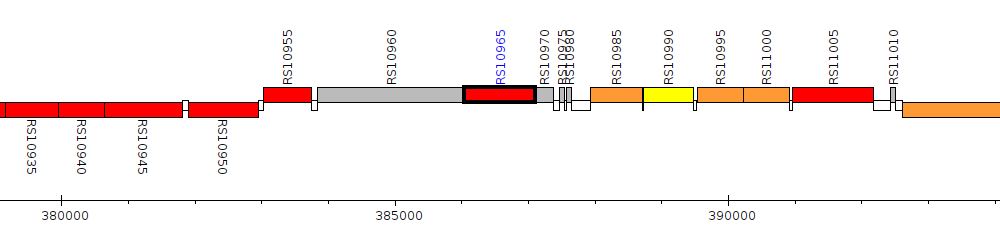
Pseudomonas tuomuerensis JCM 14085, PT85_RS10965
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006281 | DNA repair |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48150
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016798 | hydrolase activity, acting on glycosyl bonds |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR42944
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0019104 | DNA N-glycosylase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01084
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48150
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006284 | base-excision repair |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR42944
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00633
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF48150 | DNA-glycosylase | IPR011257 | DNA glycosylase | 6 | 224 | 6.12E-76 |
| Pfam | PF14815 | NUDIX domain | IPR029119 | Adenine DNA glycosylase, C-terminal | 237 | 343 | 6.6E-21 |
| FunFam | G3DSA:1.10.340.30:FF:000002 | Adenine DNA glycosylase | - | - | 21 | 135 | 1.6E-49 |
| Pfam | PF00730 | HhH-GPD superfamily base excision DNA repair protein | IPR003265 | HhH-GPD domain | 35 | 166 | 2.0E-19 |
| NCBIfam | TIGR01084 | JCVI: A/G-specific adenine glycosylase | IPR005760 | A/G-specific adenine glycosylase MutY | 6 | 274 | 2.2E-109 |
| PANTHER | PTHR42944 | ADENINE DNA GLYCOSYLASE | IPR044298 | Adenine/Thymine-DNA glycosylase | 8 | 346 | 7.3E-102 |
| CDD | cd03431 | DNA_Glycosylase_C | IPR029119 | Adenine DNA glycosylase, C-terminal | 237 | 345 | 1.61784E-15 |
| SUPERFAMILY | SSF55811 | Nudix | IPR015797 | NUDIX hydrolase-like domain superfamily | 220 | 345 | 1.11E-23 |
| SMART | SM00478 | endo3end | IPR003265 | HhH-GPD domain | 39 | 190 | 6.1E-50 |
| Gene3D | G3DSA:1.10.1670.10 | - | IPR023170 | Helix-hairpin-helix, base-excision DNA repair, C-terminal | 10 | 210 | 2.4E-88 |
| Gene3D | G3DSA:3.90.79.10 | Nucleoside Triphosphate Pyrophosphohydrolase | - | - | 225 | 349 | 6.9E-14 |
| CDD | cd00056 | ENDO3c | IPR003265 | HhH-GPD domain | 31 | 188 | 8.12447E-57 |
| Gene3D | G3DSA:1.10.340.30 | Hypothetical protein; domain 2 | - | - | 21 | 132 | 2.4E-88 |
| Pfam | PF00633 | Helix-hairpin-helix motif | IPR000445 | Helix-hairpin-helix motif | 99 | 127 | 3.9E-9 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.